













| General Information: |
|
| Name(s) found: |
gi|30265133
[NCBI NR]
gi|30259810 [NCBI NR] |
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Bacillus anthracis str. Ames |
| Length: | 808 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
RNA processing
[ISS |
| Molecular Function: |
ribonuclease R activity
[ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MEEIIQEHID KLLLFMREEA YKPLTIQELE EAFGIEGSEG FKDFVKALVT MEEKGLVIRT 60
61 RSNRYGLPEK MNLIRGKLIG HARGFAFVVP DEKKTGDDDL FIPPTELNGA LHGDTVLARL 120
121 SSQSSGSRQE GSIVRILERG TKELVGTYTE SKNFGFVIPD NKRWTSDIFV LKSASMGAVE 180
181 GHKVVVKITS YPENRLSAEG EVIQILGHKN DPGVDILSVI HKHHLPLAFP EEVMEHANSV 240
241 PETISEEDLK DRRDLRDQMI VTIDGADAKD LDDAVTVTKL ENGNYKLGVH IADVSHYVQE 300
301 GSPIDVEAAE RATSVYLVDR VIPMIPHRLS NGICSLNPKV DRLTLSCEME INNLGDVVKH 360
361 EIFQSVIKTT ERMTYADVRS ILEDEDEELM KRYEPLVPMF KEMGQLAQIL REKRMRRGAI 420
421 DFDFKEAKVL VDEEGKPTDV VMRDRSVSEK LIEEFMLVAN ETVAEHFHWM NVPFMYRVHE 480
481 DPKEDKLERF FEFVTNFGYA VKGRANEVHP RALQQILEMV QGQPEEVVIS TVMLRSMKQA 540
541 RYDADSLGHF GLSTEFYTHF TSPIRRYPDT IVHRLIREYI INGKVDNETQ AKWREKLPEI 600
601 AEHSSNMERR AVEAERETDE LKKAEYMLDK IGEEYDGMIS SVTNFGLFVE LPNTIEGLVH 660
661 VSYLTDDYYR YDEQHFAMIG ERTGNVFRIG DEITIRVINV NKDERAIDFE IVGMKGTPRR 720
721 KFKDRPVVIE QPRTGRKKRG GRSERSNERG GERGTGRKFD RGGKGKGRGS ASASTSASQP 780
781 GKKDGNGKKK KAFFENVPGF KKKKKKRK |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
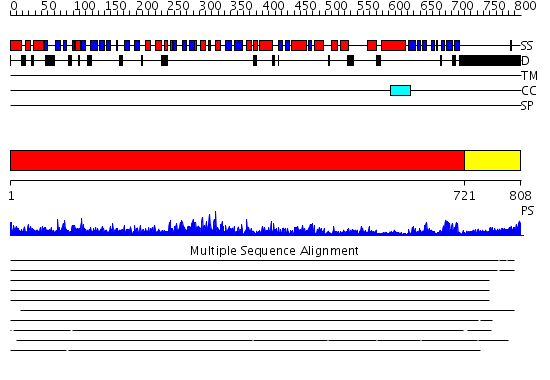
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..720] | 111.0 | No description for 2ix0A was found. | |
| 2 | View Details | [721..808] | 1.433996 | View MSA. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)