













| General Information: |
|
| Name(s) found: |
gi|50909549
[NCBI NR]
gi|46391020 [NCBI NR] gi|222623070 [NCBI NR] gi|215706900 [NCBI NR] gi|115446743 [NCBI NR] gi|113536682 [NCBI NR] |
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Oryza sativa Japonica Group |
| Length: | 652 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MQRQGRSLQR SGSKRVLDPT GGGGGDDDDH AAKRPRVPAL ASVIVEALKV DSLQKLCSSL 60
61 EPILRRVVSE EVERALAKLG PARIQGRSSP KRIEGPDGRN LQLKFTTRLS LPLFTGGKVE 120
121 GEQGAAIHVV LLDANTGVAV TSGPESCAKL DVLVLEGDFN NEEDEDWTEE EFESHIVKER 180
181 EGKRPLLTGD LQVTLKEGVG TIGELIFTDN SSWIRSRKFR LGLRVAPGSF EGIRVREAKT 240
241 EAFTVKDHRG ELYKKHYPPA LKDDVWRLEK IGKDGAFHKK LNASGIYTVE DFLQLLVKDQ 300
301 QRLRSILGSG MSNKMWESLV EHAKTCVLSG KHYVYYAIDS RNVGAIFNNI YEFTGLIADD 360
361 QFISAENLTD NQKIYADGLV KKAYEDWMHV VEYDGKALLS FKQKKKSVTT RSDTAAAATN 420
421 SPVSYGSSNT HKQLSQPAKA GQTSTGTTSE DGSTSAYNGN QAGRYAVNSQ SIPANVTTQY 480
481 ERSSLTPESQ FNGSALQNQV SRGSNILALG PPQQQHQQNF EFSALGGQSM QPTGLNPFDD 540
541 WSQPQENRSG VDDYLMEEIR MRSHEILENE EMQQMLRILS MGGASTNLTE DGFAFPNYMP 600
601 STPPNFNFGD DRARPPGKAV VGWLKIKAAM RWGIFVRKKA AERRAQLVEL ED |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
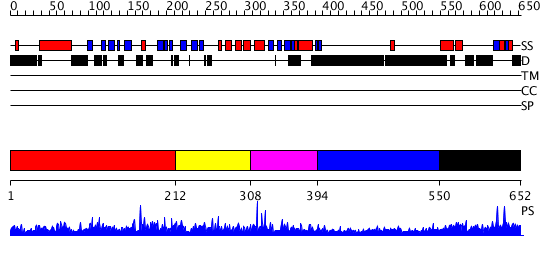
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..211] | 1.041959 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [212..307] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [308..393] | 1.070977 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [394..549] | 11.082997 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [550..652] | 1.024977 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)