













| General Information: |
|
| Name(s) found: |
SPCC1450.12
[Sanger Pombe]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 821 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA cytosol [IDA extrinsic to membrane [IEA] |
| Biological Process: |
biological_process
[ND]
|
| Molecular Function: |
phosphoinositide binding
[ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MDTASPEEID ALKRHYLKKE LFSLLIADEL NFVSEPTNLD HLGSPFVEKG KSVIPYEKSQ 60
61 IPVLRFIFKR FILTFPFLDP DSQNQLWNVN FRNLLKSLSK SNVSVLPDDS DATHLHKWLL 120
121 KFQNMLTLLM SRAFHSIQDE KNINIELLDT PKLEKDLHKS ELQQKPNILE AYVLGVRQST 180
181 QAKMLRTKRV YEYLIKVSYE ENTFFIARKY SDFSHFHHLL LKSYPNAYVP RLPPKDDHDT 240
241 YLNSSEDSTL SPLPSRSSDT NDPQSDSQHV LRQERMRIEL RQYIKNILND DELHLTEEVL 300
301 SFLTDDPVTL TASELTDINF RQELDVIRQL EQQKFVEIAS QRAKELDVYM EEFKRSLTEN 360
361 NGFTTLFTEL KEKNSISELS PSLQKTIEWA RVNLASTIYD TFIGKEKSLE TYLQIKKMHQ 420
421 LFPYTVIKNI MRFSNPLSVM KRILDTLLAQ PFGMKSLFQR LLSISLNENV RAIHKLISRY 480
481 EARIVSPEIL TKIQEQVENP CKAAREVLEK KQMKRHDYLY FILISDDVPP KLPDNLIRRV 540
541 YAERTAWKAA LDSEDYPTDP TVIRRSKRYG YMMKLMHLYA KQFNKRRSIS LISEGATGEI 600
601 MKSMVDSPEL NPNILVEQFI DLVRRHEDSF YDFVHRVYLH DSGLFASLME WIENIIGFLR 660
661 QGTSAPIDMD LVVDALDEES RQALDVELNK LLKWNQRKKS ALFLRKTTKY IPGEETSLPV 720
721 TMDELGIDEE IMAELRGDED DQDENDQVTK VEEEHMEDDD SVEEFDPIIE ERQRMKRKAN 780
781 RPANIIPPKP KLRTVKTLLP AFKEQVYPIL HKFVSENEKD I |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
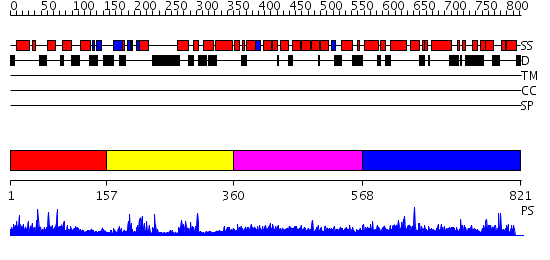
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..156] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [157..359] | 13.69897 | No description for 2dybA was found. | |
| 3 | View Details | [360..567] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [568..821] | N/A | No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)