













| General Information: |
|
| Name(s) found: |
ARFN_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 605 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IEA]
|
| Biological Process: |
regulation of transcription
[TAS]
|
| Molecular Function: |
transcription factor activity
[ISS]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MESGNVVNTQ PELSGIIDGS KSYMYEQLWK LCAGPLCDIP KLGEKVYYFP QGHIELVEAS 60
61 TREELNELQP ICDFPSKLQC RVIAIQLKVE NNSDETYAEI TLMPDTTQVV IPTQNQNQFR 120
121 PLVNSFTKVL TASDTSVHGG FSVPKKHAIE CLPPLDMSQP LPTQEILAID LHGNQWRFRH 180
181 IYRGTAQRHL LTIGWNAFTT SKKLVEGDVI VFVRGETGEL RVGIRRAGHQ QGNIPSSIVS 240
241 IESMRHGIIA SAKHAFDNQC MFIVVYKPRS SQFIVSYDKF LDVVNNKFNV GSRFTMRFEG 300
301 DDFSERRSFG TIIGVSDFSP HWKCSEWRSL EVQWDEFASF PRPNQVSPWD IEHLTPWSNV 360
361 SRSSFLKNKR SREVNEIGSS SSHLLPPTLT QGQEIGQQSM ATPMNISLRY RDITEDAMTP 420
421 SRLLMSYPVQ PMAKLNYNNV VTPIEENITT NAVASFRLFG VSLATPSVIK DPVEQIGLEI 480
481 SRLTQEKKFG QSQILRSPTE IQSKQFSSTR TCTKVQMQGV TIGRAVDLSV LNGYDQLILE 540
541 LEKLFDLKGQ LQARNQWEIA FTNNEEDKML VGEDPWPEFC NMVKKIFIYS KEEVKNLKSR 600
601 KSLSS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
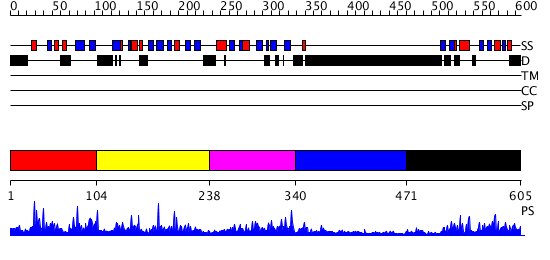
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..103] | 1.077982 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [104..237] | 35.154902 | Solution Structure of the B3 DNA-Binding Domain of RAV1 | |
| 3 | View Details | [238..339] | 4.10999 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [340..470] | 1.113984 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [471..605] | 1.21 | Solution structure of RSGI RUH-024, a PB1 domain in human cDNA, KIAA0049 |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)