













| General Information: |
|
| Name(s) found: |
TFB1A_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 591 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cellular_component
[ND]
|
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAGGQIEKLV KYKSTVKDPG TPGFLRIREG MLLFVPNDPK SDSKLKVLTQ NIKSQKYTKE 60
61 GSNKPPWLNL TNKQAKSHIF EFENYPDMHA CRDFITKALA KCELEPNKSV VSTSSEQLSI 120
121 KELELRFKLL RENSELQRLH KQFVESKVLT EDEFWATRKK LLGKDSIRKS KQQLGLKSMM 180
181 VSGIKPSTDG RTNRVTFNLT PEIIFQIFAE KPAVRQAFIN YVPSKMTEKD FWTKYFRAEY 240
241 LYSTKNTAVA AAEAAEDEEL AVFLKPDEIL ARETRHKIRR VDPTLDMEAD QGDDYTHLMD 300
301 HGIQRDGTMD VVEPQNDQFK RSLLQDLNRH AAVVLEGRSI DVESEDTRIV AEALTRVKQV 360
361 SKADGETTKD ANQERLERMS RVAGMEDLQA PQNFPLAPLS IKDPRDYFES QQGNVLNVPR 420
421 GAKGLKRNVH EAYGLLKESI LEIRATGLSD PLIKPEVSFE VFSSLTRTIA TAKNINGKNP 480
481 RESFLDRLPK STKDEVLHHW TSIQELLKHF WSSYPITTTY LHTKVGKLKD AMSNTYSKLE 540
541 AMKESVQSDL RHQVSLLVRP MQQALDAAFH HYEVDLQRRT AKSGERPNGY V |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
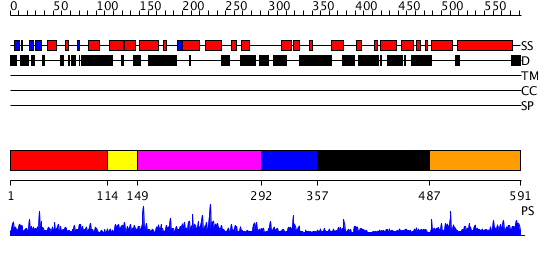
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..113] | 24.09691 | Solution structure of the N-terminal PH/PTB domain of the TFIIH P62 subunit | |
| 2 | View Details | [114..148] | 11.09691 | No description for 2diiA was found. | |
| 3 | View Details | [149..291] | 1.78 | Solution structure of the BSD domain of human Synapse associated protein 1 | |
| 4 | View Details | [292..356] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [357..486] | 1.04299 | View MSA. No confident structure predictions are available. | |
| 6 | View Details | [487..591] | 2.042989 | View MSA. Confident ab initio structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)