













| General Information: |
|
| Name(s) found: |
gi|220952606
[NCBI NR]
gi|220945978 [NCBI NR] |
| Description(s) found:
Found 2 descriptions. SHOW ALL |
|
| Organism: | synthetic construct |
| Length: | 725 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MGSQNSDDDS GSSGSSRSGS RSVTPQGGSA PGSQRSRRSG SGSDRSRSGS RSSRSRSGSG 60
61 SPRSARSGSA ESRHSQLSGS ARSKRSRSAH SRRSGSARSR KSGTPESPQS HRSGSLQSRK 120
121 SGSPQSRRSG SPQSRKSGST HSRRSGSAHS RRSGSARSRK SGSAQSDRSE SRSRSHSGSL 180
181 KGNEESRSNS PNLQIDVERA NSKSGSRSRS RSRSGSRTSR SRSKTGTPSP NRSRSGSASG 240
241 SGSDVGVPKK KARKASGSDQ EKKKSGSDSD IEESPTKAKK SRLIDTDSDS NQDVGKKAPA 300
301 AADIFGDADD ISDDEDEAGP AARKSPVRSK SRSQSKSHSH SRSMSHSRSR SRSRSRDKVE 360
361 SQVESAPKED EPEPLPETRI DVEIPRISAD LGKEQHFIKL PNFLSVVTHP FDPETYEDEI 420
421 DEEETMDEEG RQRIKLKVSN TIRWREYMNN KGDMVRESNA RFVRWSDGSM SLHLGNEIFD 480
481 AYRQPLLGDH NHLFVRQGTG LQGQSVFRTK LTFRPHSTES FTHKKMTMSL ADRSSKTSGI 540
541 KILTQVGKDP TTDRPTQLRE EEAKLRQAMR NQHKSLPKKK KPGAGEPLIG GGTSSYQHDE 600
601 GSDDENAISL SAIKNRYKKG SGAGQAEVKA STIYSSDEDE GSDFEARRSK KVDKAKASKA 660
661 LRDSDSESDA GSAKSGHSNK SGGEGGSASG SENEGSQKSG GGSSKSASGS GSGSGSGSGS 720
721 GSDND |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
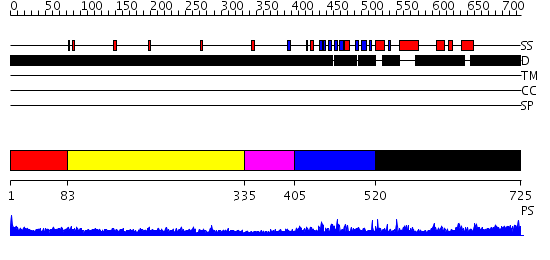
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..82] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [83..334] | 1.11199 | View MSA. No confident structure predictions are available. | |
| 3 | View Details | [335..404] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [405..519] | 1.076966 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [520..725] | 2.052985 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)