













| General Information: |
|
| Name(s) found: |
CG4673-PA /
FBpp0084267
[FlyBase]
|
| Description(s) found:
Found 18 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 652 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nuclear pore
[ISS]
|
| Biological Process: | NONE FOUND |
| Molecular Function: |
zinc ion binding
[IEA]
structural constituent of nuclear pore [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MACAPLLLEQ FIYKKRNFAY AFVRHRLFVL CKQSLIRVQS AEGIKRIEIS PKSNLKHLYD 60
61 SVQNALKVDG FGLFKERNFL TELQASGSQL VGTSLRHGDM VYLKQMAGTS SRRTSTTVLD 120
121 SQAFKTSTIS NPNSARPSFN VIEDDVDQAL SKADGTIKRE RDSKLCHHNA NGRCVHCSAL 180
181 EPYDESYLKE HNIKHLSFHS YIRKQTSGMD QGKYFVFDDI NCRIKPGCRE HPPWPKGICS 240
241 KCQPSAITLN RQTYRHVDNV MFENTKIVER FLNYWRTTGH QRMGYLYGTY EQHTDVPLGI 300
301 RAKVAAIYEP PQESTRDSIN IQPDEFADDV DAVASALGLK KIGWIFTDLI TDDASIGTVK 360
361 QIRGIESHFI TAQECITAGE LQNRHPNPCK YASNGVFGSK FVTICVTGDK TKQVHMEGYA 420
421 VSAQCMALVR DNCLIPTKDA PELGYVREST DKQYVPDVFY KEKDLYGNEV QRLARPLPVE 480
481 YLLVDVPAST PLQPIYTFTE YDKRQPFPIE NRYIDGHLQD FNALSCYLSA WGEEEFLEAI 540
541 SDFHLLVYLY KMDMLPLRQH MGPLLEAVRT KNPNQAAQFK VVDVWKLLES LIQASSGGSG 600
601 GTTSYPSGGA SASAGASGSE AMDLDANTWT CNHCTFINRG ELTSCEICSL PR |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
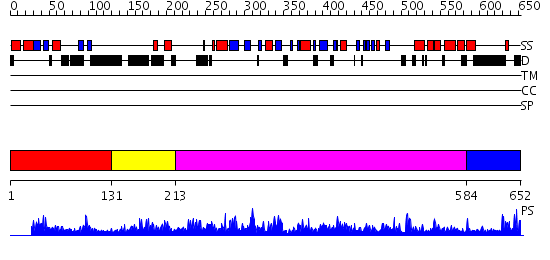
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..130] | 14.30103 | Solution structure of a novel beta-grasp fold like domain of Hypothetical protein (Arabidopsis thaliana) | |
| 2 | View Details | [131..212] | 42.19382 | No description for PF05020.6 was found. No confident structure predictions are available. | |
| 3 | View Details | [213..583] | 2.8 | No description for 2og4A was found. | |
| 4 | View Details | [584..652] | 10.041995 | View MSA. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)