













| General Information: |
|
| Name(s) found: |
MCPH1-PA /
FBpp0290643
[FlyBase]
|
| Description(s) found:
Found 20 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 779 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA]
|
| Biological Process: |
embryonic development
[IMP]
mushroom body development [IMP] pole cell formation [IMP] mitosis [IMP] |
| Molecular Function: |
molecular_function
[ND]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSTNEPSITN KLRSLLLDDQ AARDRLMLKP ALAVSKAISQ HRSEFHQDEN EAGAEGSAAK 60
61 PDPPIGAATP TRQEGHWARL QQDLNSPNAA LRLRAIRALK SPTKSAYNTF DVPLAEQSII 120
121 TMEERNPQEP PSLPELLKDI VVYVEVRTGN DNRSEGVKSI IAKMGAQVKD RLTRTTTHVV 180
181 FKDGLLSTYK KATEWNIPVV SILWIEACKV QLKICDPKQF PISNIRMYEY PELFGKMPRV 240
241 KFMQPDSELN QRPRKRQGTP TSSKDNEPGS TNKRAQTGTP TTTTPKNDIS RFFRAINHNK 300
301 PSDDEAIESP ATKLLNRISS GCYTPLPAGG TPQLMGTNGS GDAVAQKVQA QKSLQFGEQT 360
361 DSNTPQAKPS STSRPRRSTV DPPPVMTPPV RTTRRRSCVA EISSPAATDT RMTRRRSSLL 420
421 TSTVIHEADP TQTAVAEPRM TRRRSSLLGI AKQAEETAQR QTIQPITEEL APVEENRNSI 480
481 IMAETNIINN TNCLEQSIEI TAVEAEASFK ADRRGTLYAS ELMDISSPRR RSVAKSSLEL 540
541 RQENLAPRAR QSLSNVTTQT SFLDVEQQVT PLFSSTRLPT GQTTGNRRRT IFNMDMDVIN 600
601 ESIERMNQSH RMSLALANKA ELGESCQVHE QEPPQAEPVP SQEPKRRRLF NPNDEVVISP 660
661 AMSNRKSRLS ITGSGSSSSV QKRRRSLATP AKSSSSELCA TTPLSKAVTN KEAEKSKSNT 720
721 EISEASLDAV PEVKKSTSNA GEENEKPKGK SRTRIRTLVH TNMHQEQINV IRKVGNTTE |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
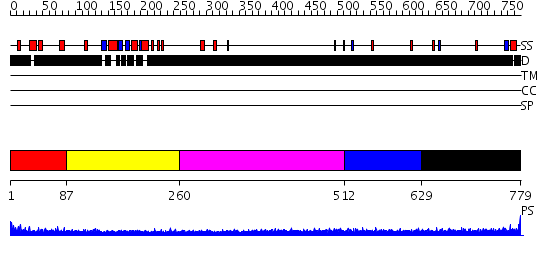
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..86] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [87..259] | 4.46 | No description for 2d8mA was found. | |
| 3 | View Details | [260..511] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [512..628] | 3.202988 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [629..779] | N/A | No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)