













| General Information: |
|
| Name(s) found: |
Dyb-PC /
FBpp0087041
[FlyBase]
|
| Description(s) found:
Found 19 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 623 amino acids |
Gene Ontology: |
|
| Cellular Component: |
dystrophin-associated glycoprotein complex
[ISS]
dystrobrevin complex [ISS][NAS] |
| Biological Process: | NONE FOUND |
| Molecular Function: |
zinc ion binding
[IEA]
structural constituent of muscle [ISS] calcium ion binding [IEA] cytoskeletal protein binding [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MLEVVFGRGM ELEPRVAILQ DLRLQTFDSI RFASYRTASK LRYIQKSTNL HLVDIWNVIE 60
61 AFRENGLNTL EPQSEVSVAR LETLVSSLYH NLNKRLPTAQ QVPVDSKAGL LLNWLLAAYT 120
121 SDNSGKIRVF SIKVALATMC SGKLVDKLRY IFSQISDGAG QLVAWKLGEF LREVLALPAA 180
181 VYESPTFHYK EGLEEEIFPA ENKVTVNDFM ATLMSEPGPS CLVWLPLVHR LATVETIVHP 240
241 TVCSVCHKEN FTGFRYRCQR CHAYQLCQEC FWHGKTSLNH QNDHEVKEYS SYKSPSKQIG 300
301 HSLRKSFRCV PEKTVQVLPR FPDQPEKTLN LSHIVPPSPL PSHNGFSDPS GLVHGHHGPH 360
361 PGLPGQHGLF DRSSTLDSRA TGRSLDSASG TAGTTMSRVA AASANDEEHR LIARYAARLA 420
421 QENRAPSNLP DNATPIGTDN SRAQRELIAQ LESKNKEIMR EIARLRRQQE TEQMAPENPA 480
481 LINELRALRQ RKGELEGHLG ALQDSRRQLM EQLEGLMRML KNQQTASPRS TPNSSPRSGK 540
541 SPPMPGGAGI LGTSPMSALH AQQMQQHPMA GGVRASQQQL LQQQQQQAHG HGPFSQSQLE 600
601 QLNQISSDMR SAFAANGSAS KCE |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
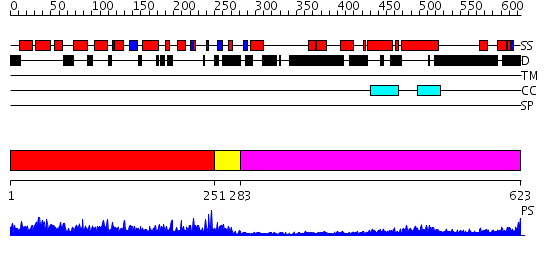
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..250] | 85.154902 | Dystrophin | |
| 2 | View Details | [251..282] | 3.08 | No description for 2e5rA was found. | |
| 3 | View Details | [283..623] | 9.522879 | No description for 2dfsA was found. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.94 |
Source: Reynolds et al. (2008)